The Endpoint
Carcinogenicity
The process of carcinogenesis involves the transition of normal cells into cancer cells via a
sequence of stages that entail both genetic alterations (i.e. mutations) and non-genetic events.
Genotoxic modes of action involve genetic alterations caused by the chemical interacting directly with DNA to result in a change in the primary sequence of DNA.
Non-genotoxic modes of action include epigenetic changes, i.e., effects that do not involve alterations in DNA but that may influence gene expression, altered cell-cell communication, or other factors involved in the carcinogenic process.
Carcinogens
Chemicals are defined as carcinogenic if they induce tumours, increase tumour incidence and/or malignancy or shorten the time to tumour occurrence.
Chemicals can induce cancer by any route of exposure (e.g., when inhaled, ingested, applied to the skin or injected), but carcinogenic potential and potency may depend on the conditions of exposure (e.g., route, level, pattern and duration of exposure). Carcinogenic chemicals have conventionally been divided into two categories according to the presumed mode of action: genotoxic or non-genotoxic.
Information Requirement
For the endpoint of carcinogenicity, standard information requirements are specifically described for substances produced or imported in quantities of ≥1000 tons/year.
The precise information requirements will differ from substance to substance, according to the toxicity information already available and details of use and human exposure for the substance in question.
A carcinogenicity study may be proposed by the registrant or may be required by the Agency if:
- the substance has a widespread dispersive use or there is evidence of frequent or long-term human exposure; and
- the substance is classified as mutagen category 3 or there is evidence from the repeated dose study(ies) that the substance is able to induce hyperplasia and/or pre-neoplastic lesions.
If the substance is classified as mutagen category 1 or 2, the default presumption would be that a genotoxic mechanism for carcinogenicity is likely. In these cases, a carcinogenicity test will normally not be required.
Carcinogenic potential
The objective of investigating the carcinogenicity of chemicals is to identify potential human carcinogens, their mode(s) of action, and their potency. The golden standard test under REACH for carcinogenicity is OECD 451. OECD guideline no. 451 recommends the use of 400 animals (rats and mice) for the studies.
Hazard assessment
Carcinogenic potential may be identified from epidemiological studies, from animal experiments and/or other appropriate means that may include (Quantitative) Structure-Activity Relationships ((Q)SAR) analyses and/or extrapolation from structurally similar substances (read-across).
Once a chemical has been identified as a carcinogen, there is a need to elucidate the underlying mode of action (genotoxic or non-genotoxic). Human studies are generally not available for making a distinction between the modes of action. A conclusion on this depends on the outcome of mutagenicity/genotoxicity testing and other mechanistic studies. In addition to this, animal studies (e.g. the carcinogenicity study, repeated dose studies, and experimental studies with initiation-promotion protocols) may also inform on the underlying mode of carcinogenic action.
For genotoxic carcinogens exhibiting direct interaction with DNA, it is not generally possible to infer the position of the threshold from the no-observed-effect level on a dose-response curve, even though a biological threshold below which cancer is not induced may exist. For non-genotoxic carcinogens, no-effect-thresholds are assumed to exist and to be discernable.
The Model
To address the carcinogenicity issue two complementary approaches were investigated:
- a regression approach
- a continuous approach
The original dataset used for the development of CAESAR model for carcinogenicity contains 805 chemicals extracted from CPDBAS with associated TD50 values for rat. In classification any compound with a finite TD50 dose was associated with the toxic class, while non positive compounds were assigned to the non toxic class. This dataset was then spit into training (n = 644) and test (n = 161) sets. In regression, only compounds with a TD50 dose were used.
Regression Model
In collaboration with the EC project CHEMPREDICT we developed quantitative structure - activity relationships (QSAR) models based on SMILES. Simplified molecular input line entry system (SMILES) has been used as elucidation of the molecular structure for QSAR to predict carcinogenicity. Using the Monte Carlo method we constructed optimal descriptors, which are a mathematical function of composition of the SMILES elements together with special codes of cycles present in molecules.
Good results have been obtained on both the training and test set, as shown below.
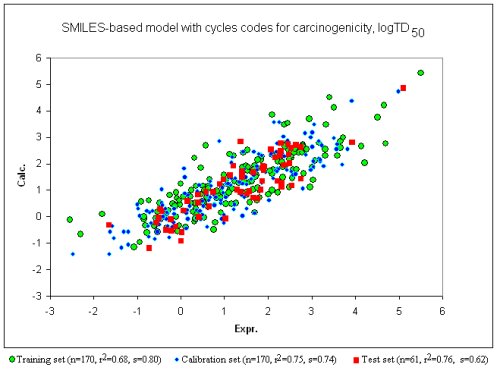
The graphic above show the correlation between the experimental TD50 (x-axis) and that calculated by the model (y-axis).
Classification Model
A classification model has been developed adopting the Counter-Propagation Artificial Neural Network (CP-ANN) method and a set of MDL Chemical Descriptors.
Results of the binary classification model are shown below:
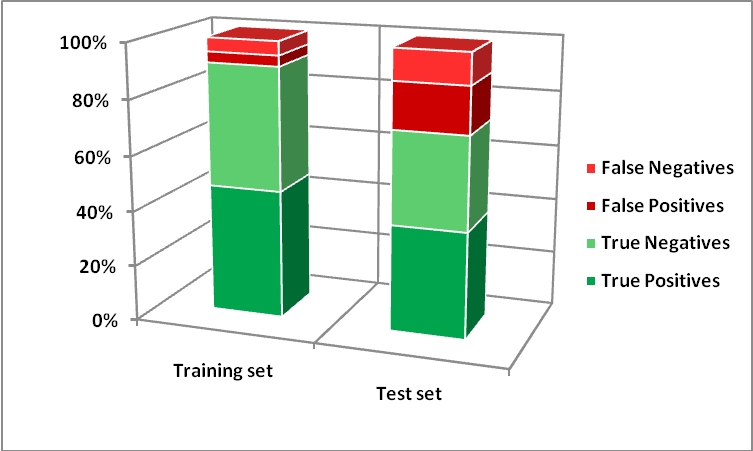
Notice for the use of in silico CAESAR models addressing human toxicology related to
human toxicology (i.e.: carcinogenicity and developmental toxicity models).
Currently, the role of in silico models in these endpoints can be limited to consider them as an
ingredient in deriving a weight of evidence rather than to substitute per se existing methods.
Their utility is consequently in support to the overall assessment.
The user is also advised that, since some of the models are based on datasets focused on a limited
chemical space, particular attention should be placed for these two endpoints in the evaluation of
similar compounds already present in the studied datasets and the model's ability to correctly
predict them.
References
- New public QSAR model for carcinogenicity.
Fjodorova N., Vračko M., Novič M., Roncaglioni A., Benfenati E.
Chemistry Central Journal 2010, 4(Suppl 1):S4 (29 July 2010)